|
| |
| Genomic Services |
|
| Protein Services |
|
| |
| |
Located in rooms
B065 and B017
|
|
PAN is offering sample prep services for Next Generation Sequencing technology. You must check the requirements for each service before submitting your samples to us.
Full Service: (back to top)
• General Info. --
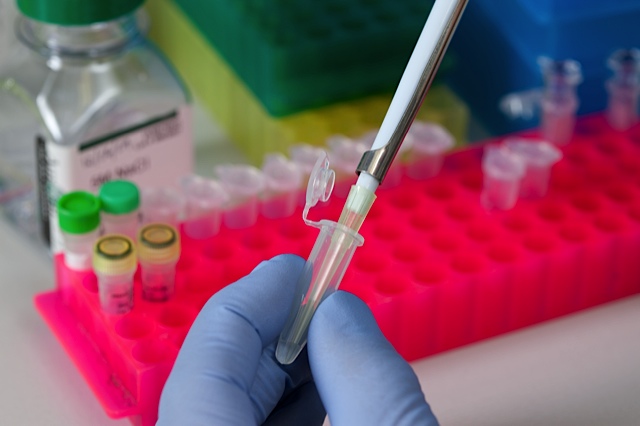 We understand that preparing high quality libraries is the very first critical step of the next generation sequencing workflow. Depending on your desired sequencing application, we will work with you to identify and use library construction methods to suit your experimental needs with the goal of generating high yield and high quality libraries. We understand that preparing high quality libraries is the very first critical step of the next generation sequencing workflow. Depending on your desired sequencing application, we will work with you to identify and use library construction methods to suit your experimental needs with the goal of generating high yield and high quality libraries. |
• Applications for Different Sequencing Platforms --
We are currently providing library preparations starting with purified DNA or RNA. We can generate libraries for different applications – see table. The libraries can be prepared, based on your requests, for different sequencing platforms, such as Illumina MiSeq, Illumina HiSeq, Ion Torrent. |
• Library Preparation Kits --
There are many different library preparation kits available commercially. The two major vendors are Illumina and New England Biolabs (NEB).
You are welcome to visit their websites and check out the different criteria for each kit and let us know which would fit best for your sample(s). If you are unsure of which kit to choose from, please contact us and we would be more than happy to work with you to determine the kit best suited for your project.
• Exome Capture --
As part of the full service, we also offer targeted exome capture, a way to isolate exonic regions of interest in the human genome or to identify coding variants. You will need to let us know the number of samples needed for pooling and the desired exome capture protocol for your project. Exome capture kits/protocols are available from Agilent, Illumina, and Nimblegen. If you are unsure of which kit to use, we can help you find the kit best suited for your project.
For targeted capture for both exome and custom regions of interest you must purchase the capture kit from PAN and we will provide the remainder of consumables and labor. |
PAN is currently supporting kits from Illumina. The following table can be used as a guide to choose an appropriate kit for your project.
(Click on the name of each kit for more detailed information)
*TruSeq LT: 24-plex; TruSeq HT: 96-plex |
• Sample QC and Library QC on MiSeq --
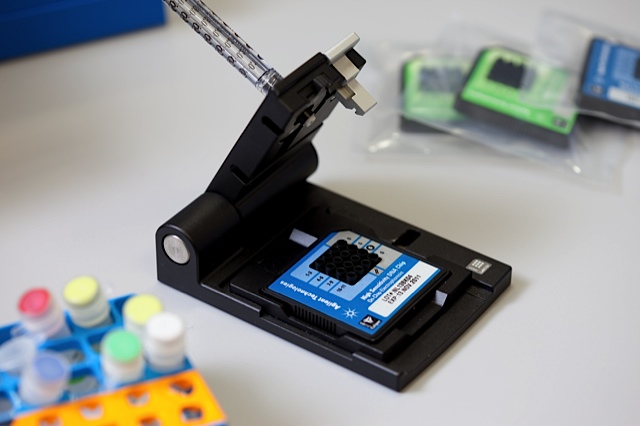 Throughout the whole library preparation process, QC steps will be performed to ensure the quality of the sample. As for library validation, QC and quantification will be done on Agilent Bioanalyzer and qPCR, respectively. Throughout the whole library preparation process, QC steps will be performed to ensure the quality of the sample. As for library validation, QC and quantification will be done on Agilent Bioanalyzer and qPCR, respectively.
For further QC of the library, you can request to have it run on our MiSeq machine. The collected data (~3GB), including the Q-Scores and cluster density, will help you determine the accuracy and quality of the prepared library..
If your sample passes QC and you decide to move forward with deep sequencing, we can transfer your samples directly to the sequencing facility of your choice. In the event that the sample(s) failed QC and you still wish to proceed with sequencing, we would require a written confirmation (email will suffice) stating that you wish to continue with the sequencing experiment and will pay for all requested services regardless of the outcome. If your sample is low concentration and you submit additional material, we may require an additional round of QC to ensure that the material will yield sufficient clusters and quality sequence. Please note that multiple freeze-thaw cycles of prepared libraries have been shown to degrade library performance so it is important to keep these to a minimum. |
|
|
MiSeq Run : (back to top)
• General Info. --
 We currently have an Illumina MiSeq in house. This benchtop sequencer is capable of producing high quality data at a much shorter run time compared to other sequencers. It yields highly accurate results with a high level of base calls above Q30 for a wide range of applications including targeted resequencing, small-genome sequencing, RNA sequencing, ChIP-Seq, and library QC. We currently have an Illumina MiSeq in house. This benchtop sequencer is capable of producing high quality data at a much shorter run time compared to other sequencers. It yields highly accurate results with a high level of base calls above Q30 for a wide range of applications including targeted resequencing, small-genome sequencing, RNA sequencing, ChIP-Seq, and library QC.
You can visit Illumina website to check out the detailed specifications and the different sample prep kits and run cycle kits for different applications.
The table below will give you guidance to choose the right run cycle based on the application needed.
| Application/Project Type |
Recommended Read Length |
Targeted Resequencing
| Amplicon (tens of targets) |
| Amplicon (hundreds of targets) |
| Hybrid Capture (thousands of targets) |
| 16S Metagenomics |
| Clone Checking |
|
| 1 x 250 |
| 2 x 250 |
| 2 x 75 |
| 2 x 150 |
| 1 x 36 |
|
Small Genome
| De novo |
| Resequencing |
| Plasmids |
|
|
RNA Sequencing
| Small RNA |
| RNA-Seq (microbial) |
|
|
| ChIP Seq |
|
| Library QC |
|
| ** For more information on other applications and recommended sample prep kits, please refer to this brochure and Illumina website. |
Specs for MiSeq
MiSeq Reagent Kit |
Clusters |
Read Length (BP) |
Output |
Total Time |
Q Score (>Q30) |
Standard |
~15M |
2 x 25 cycles |
~2.25 Gb |
~5.5 hrs |
> 90% |
| 2 x 150 cycles |
~4.5 Gb |
~24 hrs |
> 80% |
| 2 x 250 cycles |
~7.5 Gb |
~39 hrs |
> 75% |
| 2 x 300 cycles ** |
~15 Gb |
>48 hrs |
> 70% |
Micro |
~4M |
2 x 150 cycles |
~1.2 Gb |
- |
- |
Nano |
~1M |
2 x 150 cycles |
~300 Mb |
- |
- |
| 2 x 250 cycles |
~500Mb |
- |
- |
** Coming soon.. |
• Sample QC --
Prepared library submitted will be run on Agilent Bioanalyzer as part of the initial QC process for a MiSeq run. If your sample failed QC and you still wish to proceed with sequencing, we would require a written confirmation (email will suffice) stating that you wish to continue with the sequencing experiment and will pay for all requested services regardless of the outcome. If your sample is low concentration and you submit additional material, we may require an additional round of QC to ensure that the material will yield sufficient clusters and quality sequence. Please note that multiple freeze-thaw cycles of prepared libraries have been shown to degrade library performance so it is important to keep these to a minimum. |
• Data Release --
 The data generated from MiSeq will be accessible and downloadable online through BaseSpace, a free system offered from Illunina that allows data sharing amongst collaborators. BaseSpace also contains different analysis applications that can analyze your data and give you information about alignment, structural variants, and/or contig assemblies for each genome requested and each sample. The data generated from MiSeq will be accessible and downloadable online through BaseSpace, a free system offered from Illunina that allows data sharing amongst collaborators. BaseSpace also contains different analysis applications that can analyze your data and give you information about alignment, structural variants, and/or contig assemblies for each genome requested and each sample.
**
You'll need to sign up for an Illumina account first in order to use BaseSpace.
If you only need the raw data and use your own analysis software, you can request us to set up a link for you to download the data directly from our server. |
|
|
|




 We understand that preparing high quality libraries is the very first critical step of the next generation sequencing workflow. Depending on your desired sequencing application, we will work with you to identify and use library construction methods to suit your experimental needs with the goal of generating high yield and high quality libraries.
We understand that preparing high quality libraries is the very first critical step of the next generation sequencing workflow. Depending on your desired sequencing application, we will work with you to identify and use library construction methods to suit your experimental needs with the goal of generating high yield and high quality libraries.  Throughout the whole library preparation process, QC steps will be performed to ensure the quality of the sample. As for library validation, QC and quantification will be done on Agilent Bioanalyzer and qPCR, respectively.
Throughout the whole library preparation process, QC steps will be performed to ensure the quality of the sample. As for library validation, QC and quantification will be done on Agilent Bioanalyzer and qPCR, respectively.