
|
||||||||
| Genomic Services | ||||||||
|
||||||||
| Protein Services | ||||||||
|
||||||||
Located in rooms
B065 and B017 |
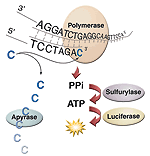
| Pyrosequencing |
• SNP/mutation and DNA methylation Analyses |
**Essential software available for download:
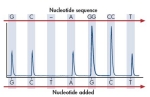
SNP/Mutation Analysis• Analyze single and multiple SNPs and insertion/deletion mutations within 200bp stretches of the genome. • Pyrosequencing technology presents SNPs in the context of the surrounding sequence. This guarantees that each individual analysis is correct, without the need for other controls. In a clinical context, this level of certainty is highly valuable, especially as the genetic methods developed in today’s research mature into tomorrow’s genetic tests.
|
||
DNA Methylation Analysis• Quantitatively analyze PCR products amplified from bisulphite treated genomic DNA for levels of methylation at single or multiple CpG sites across the genome. Typical workflow for analysis of CpG methylation • Pyro Q-CpG quantifies methylation in explicit sequence context, and is both fast and easy to perform. Assay design is flexible, since the distance from the first base to be sequenced can be varied, and therefore the primer can usually be positioned in a region free of CpG sites. In addition, there are four options for design; the assay can be performed in forward or reverse orientations on either the top or the bottom strands.
|
Home | FAQs | | Prices | Contact Us | Publications | Feedback
Beckman Center | Stanford Medical Center | Stanford University
© 2006 Stanford PAN Facility. All rights reserved
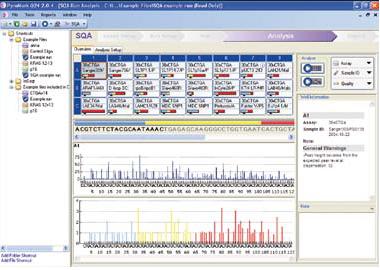
 • Results from mutation analysis of codons 12 and 13 using PyroMark KRAS v2.0 test and PyroMark Q24. The upper Pyrogram trace shows a sample with a normal genotype. The middle Pyrogram trace shows mutation analysis in a sample with a G to A mutation in position 2 of codon 12, and the lower Pyrogram trace shows a sample with a G to T mutation in position 1 of codon 12 identified post-run by the altered sequence. Light blue areas indicate the variable position. (From Qiagen)
• Results from mutation analysis of codons 12 and 13 using PyroMark KRAS v2.0 test and PyroMark Q24. The upper Pyrogram trace shows a sample with a normal genotype. The middle Pyrogram trace shows mutation analysis in a sample with a G to A mutation in position 2 of codon 12, and the lower Pyrogram trace shows a sample with a G to T mutation in position 1 of codon 12 identified post-run by the altered sequence. Light blue areas indicate the variable position. (From Qiagen)